Technology innovation for crop improvement
KeyGene is the go-to research company for the development and application of breakthrough technology innovation. We do this for and with our shareholders and other partners, for crop improvement in for instance vegetables, fruits, ornamentals, and industrial field crops.
Technology innovation
We develop and apply breakthrough technology innovation for crop improvement, for and with partners around the globe.
Crop improvement
We use proprietary technology innovation to support partners around the globe to boost their crop improvement.

9 April 2024
Advancing Smart Breeding for Resilient Agriculture: A Conversation with CropXR Managing Director Hedwich Teunissen
Hedwich Teunissen, Managing Director of CropXR, shares insights into the institute's misison, science and collaborative approach.
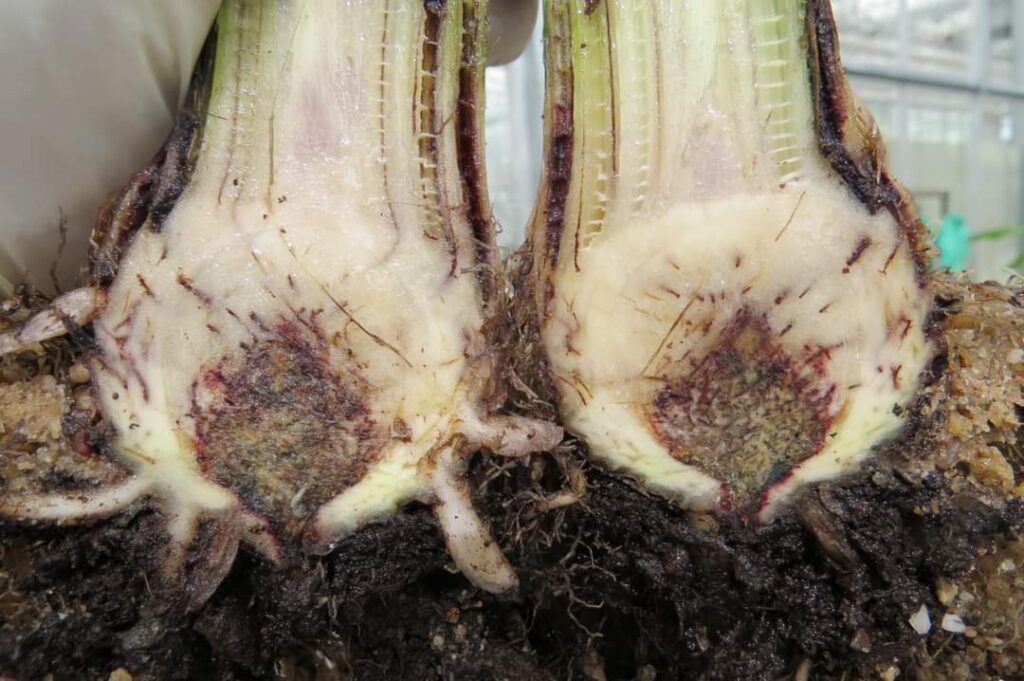

6 March 2024
Recent evolution of Fusarium TR4 strain has increased diversity in ‘virulence’-DNA of the fungus
Paper: Evolution of Fusarium TR4 strain has increased diversity in ‘virulence’-DNA of the fungus
22 February 2024
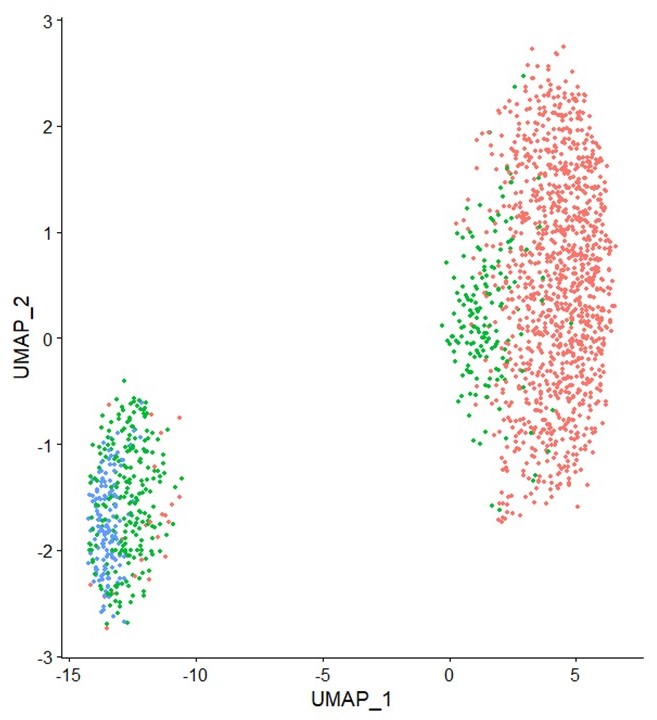
Studying nuclei of individual pollen grains is feasible now
A collaboration between Takara Bio and KeyGene has made it possible to carry out single-nucleus RNA sequencing (snRNA-seq) successfully.
Take a look at our work
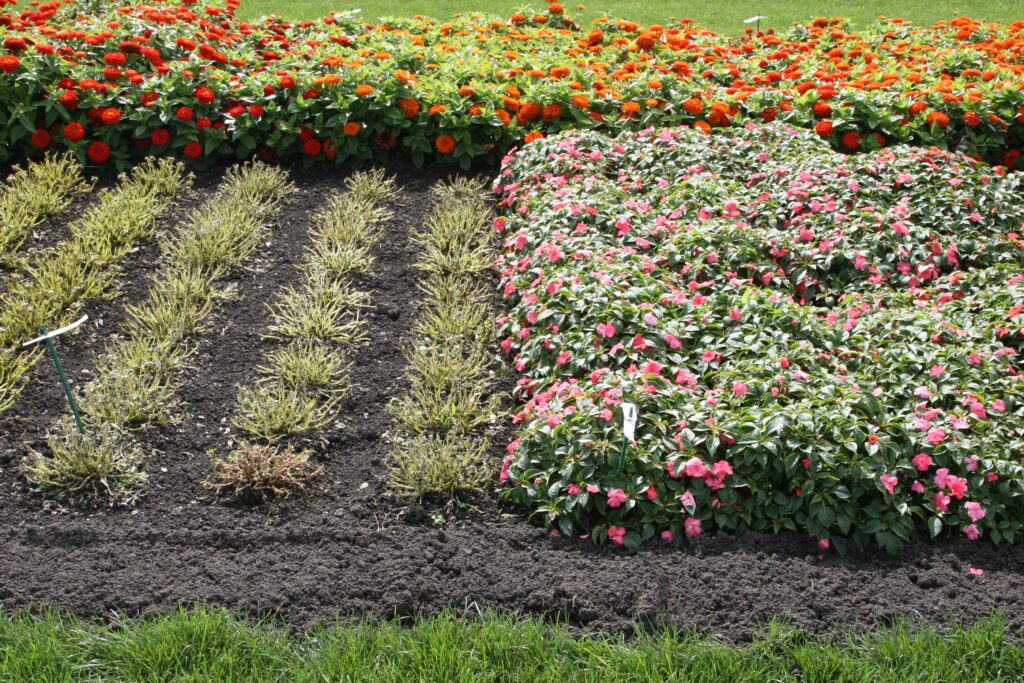

Accelerated breeding of impatiens resistant to downy mildew
KeyGene supported Ball Horticultural in the USA to reveal, understand and utilize the genetic variation at hand within their germplasm, for accelerated breeding of impatiens varieties resistant to downy mildew