Licensing KeyGene's technology innovations
We are enthusiastic about our breakthrough technology innovations. Through licensing, KeyGene aims to provide its partners access to its IP that can help to create competitive advantages for our partners.
Our technologies and trait components are accessible to breeding companies, academic organizations, as well as commercial service providers. For non-profit research organizations, favorable conditions for access apply.
Many global seed companies including Bayer, Enza Zaden, Rijk Zwaan, Limagrain Vegetable Seeds and Takii & Co. Ltd have one or more licenses for our technologies. KeyGene’s technologies also find application outside Agrofood, such as in cancer diagnostics.
Our innovations

Pollen viability
Improve fertility and viability in interspecific hyrbids, using compounds that inhibit the MRN-ATM pathway.
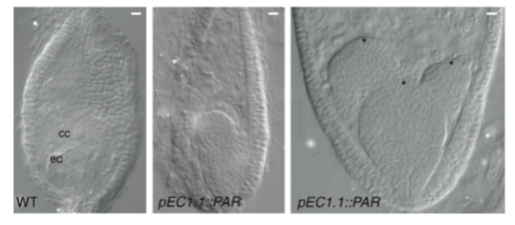
Parthenogenesis promotor
Promoter sequences that convert a sexual allele into a parthenogenesis allele, which is capable of inducing parthenogenesis.

Coregeneration of recalcitrant plants
Regenerate recalcitrant plant cells into regenerative shoots by establishing contact to regenerative plant cells.
Msi2 gene for DH induction
Mutant Musashi RNA-binding protein 2 (Msi2) proteins, nucleic acid sequences, and plants for haploid inducer lines.
CENPC Gene for DH induction
Mutant Centromere Protein C proteins, nucleic acid sequences, and plants for haploid inducer lines.

Efficient plant selection
Plant selection based on genotypical and/or phenotypical data sets and deep learning.

Target enrichment through selective sequencing
A combination of TarSeq and subsequent selective sequencing, a highly effective method for target sequence enrichment.

Covalently closing DNA strands by TelN for NGS library preparation
TelN adapters / primers for NGS library preparation for amplification free enrichment of target sequences.

Improving drought resistance: pectinesterase
For plants with increased drought resistance by impairing the expression of functional pectinesterase.
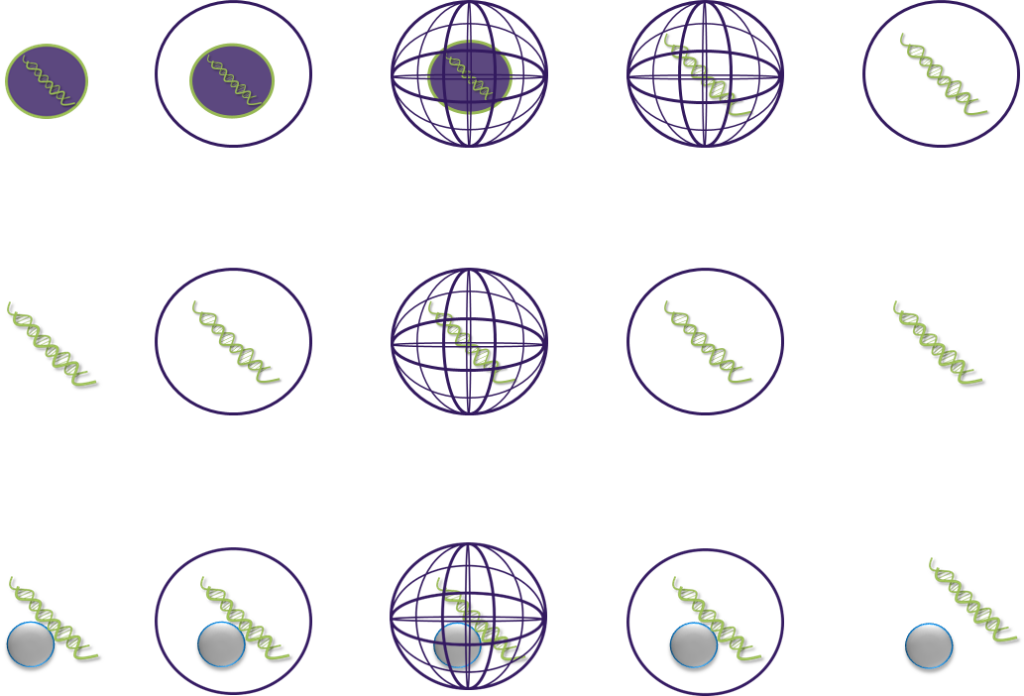
Semi-solid state for nucleic acid handling
For methods and kits for HMW nucleic acid library preparation, effectively reducing shear stress.

Targeted Sequence Addition
Method to increase shoot regeneration by overexpressing genes for histidine kinases

Shoot regeneration by CHK genes
Method to increase shoot regeneration by overexpressing genes for histidine kinases
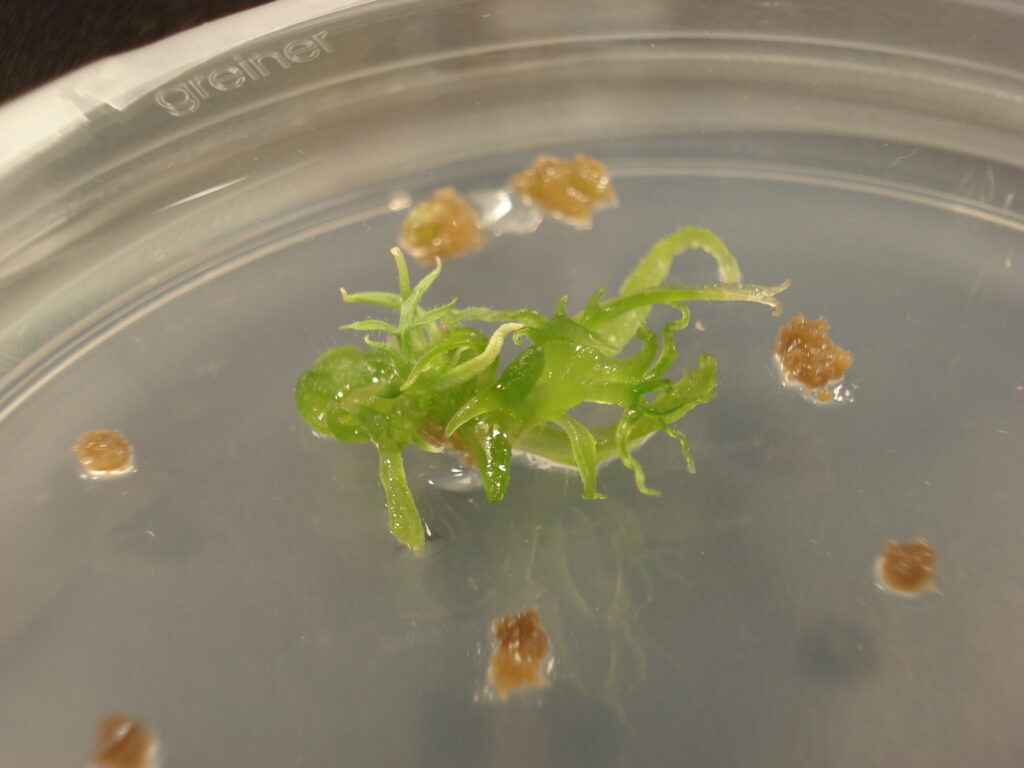

Dual guide for CRISPR editing of plant cells
Effective genome editing CRISPR system nuclease guided by non-covalently linked, crRNA and tracrRNA
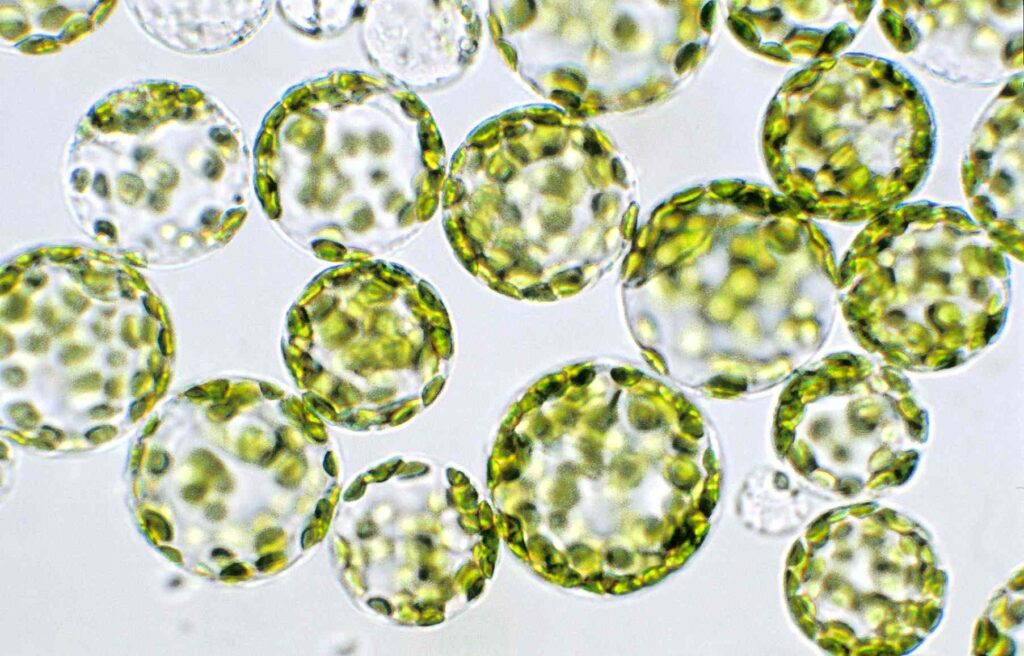
Genes for hormone free regeneration
Transient expression of 2 genes circumvents the need for cytokinin and auxin

Germacrene A Synthase mutants
Decreased sesquiterpene lactones and increased levels of squalene & phenolic compounds
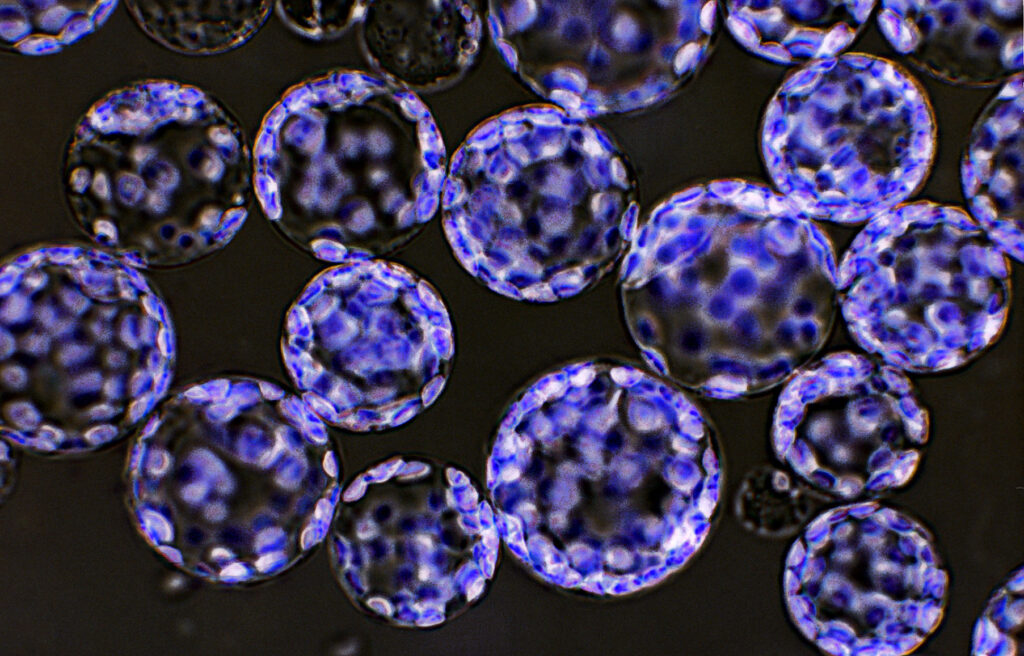
Mad7 for genome editing in plant cells
Highly effective Type V CRISPR system for plant cell genome editing
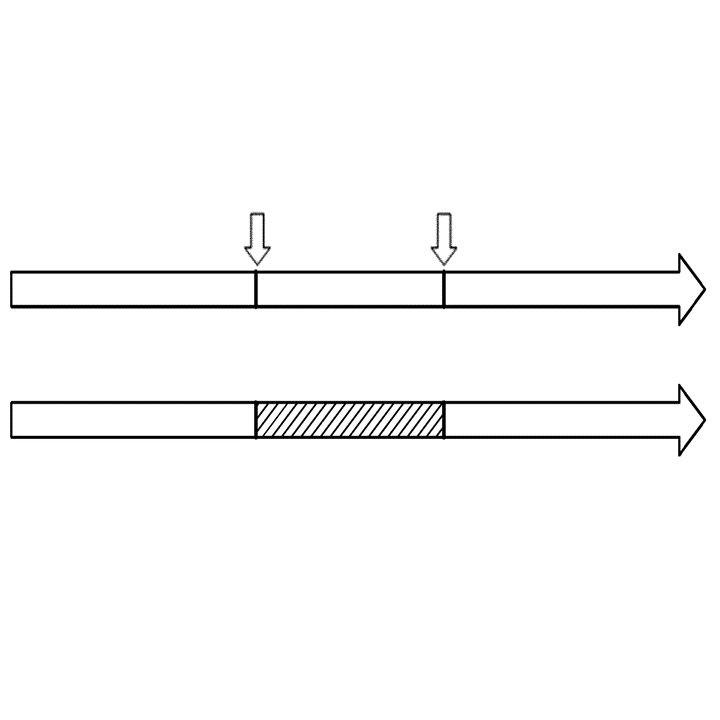
Genome editing for altered protein function
Editing of a coding sequence by two site-specific nucleases that causes a frameshift
Genome editing by programmed deaminases
Genome editing using cysteine or adenine deaminase fused to a site-specific nuclease e.g. a CAS
Glycerol-free transfection of CRISPR complex
Glycerol free genome editing of protoplasts using CRISPR-CAS protein and in vitro transcribed gRNA

Targeted translocation
Translocation between 2 homologous/homeologous chromosomes via double-strand break and rejoining
Oligonucleotide-delivery to protoplasts
Method to effectively deliver mutagenic nucleobases to plant protoplasts using polyethylene glycol
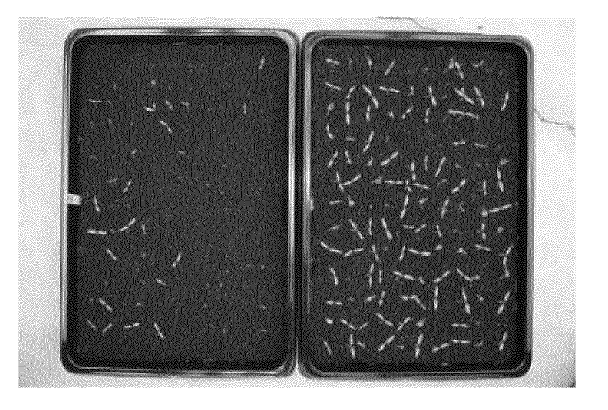
Improved seed germination
Seeds with improved germination quality, by pollination of graft hybrid mother plants.

2S1 Graft hybrid technology
Plants with improved epidermis-based complex traits, e.g. biotic stress resistance
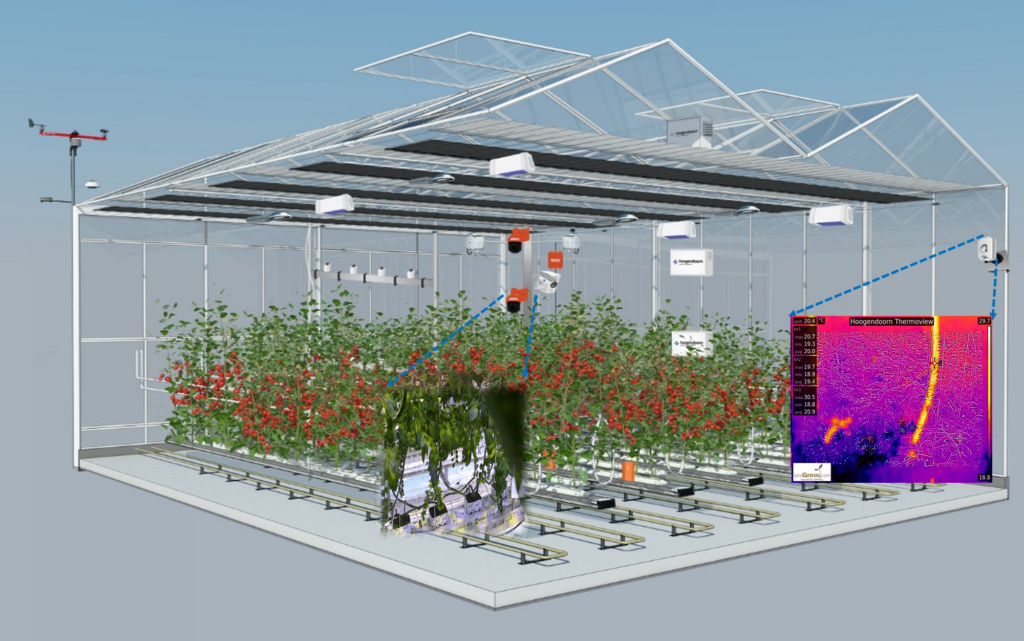

Virtual reality greenhouse
3D visualization of individual plants and association with plant characteristic data

Improving Drought Resistance in Plants: UPL3
Plants with increased drought resistance by impairing the expression of functional UPL3

Improving drought resistance in plants: UPL4
Plants with increased drought resistance by impairing the expression of functional UPL4
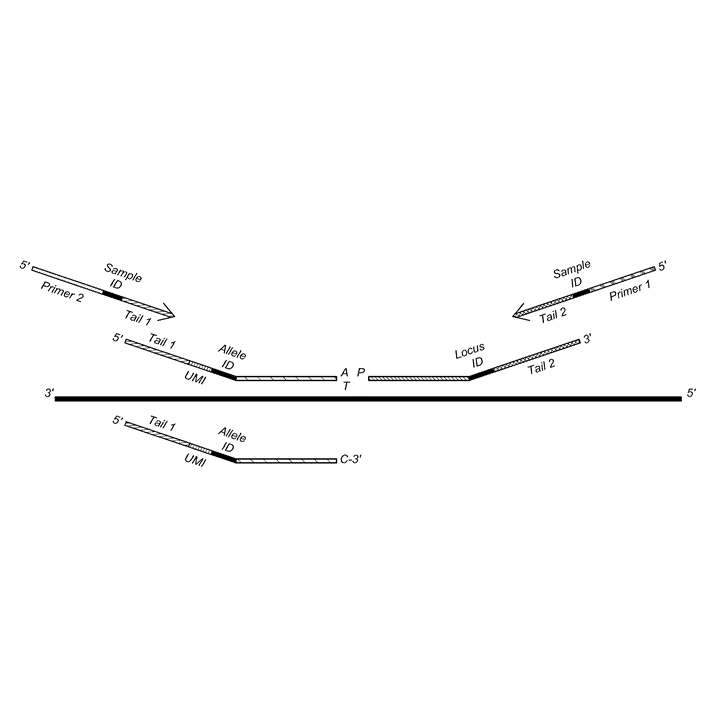
Genotyping polyploids using molecular tags
Effective genotyping of polyploids using probes comprising unique molecular indices
Accurately amplifying mixtures of probes
Production of oligonucleotides, in particular targeting oligonucleotides or nucleic acid probes.
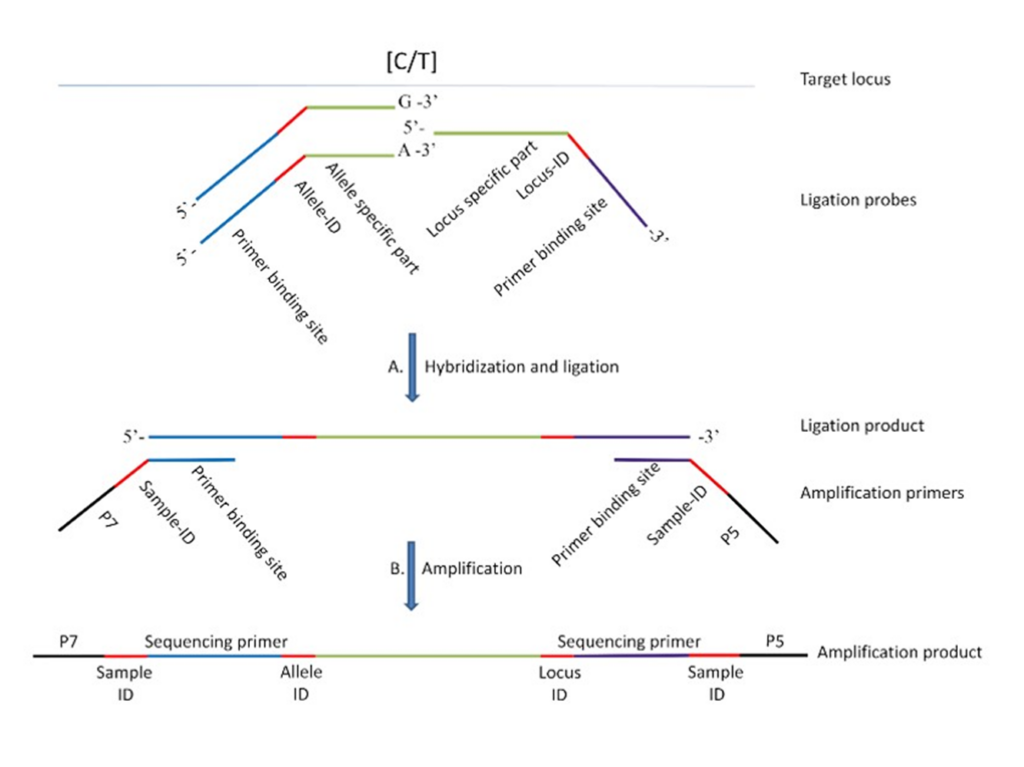
SNPSelect
Genotyping, based upon locus & allele identifiers; a way to amplify and process DNA into probes

Directed genomic selection
Genomic selection using genome-wide complementarity, taking into account recombination probabilities
Combinatorial barcodes
At least two nucleotide sequence barcodes, for identifying sample origin in NGS.
KeyPoint® Breeding
Multiplex detection of variation of genes / loci of interest, based on high throughput sequencing.